Only matches that had an e worth of 10 five or lower and had sequence similarity of 50 base pairs or greater were included in our MG RAST evaluation. Metagenomes were also analyzed that has a neighborhood BLASTN to a database of N metabolism genes that we constructed with searches at the NCBI internet site. The database included the known genes for your enzymes concerned in denitrification, DNRA, and Annamox, as these processes are nitrate reduction pathways. The extremely profiled functional genes for nitrifi cation and nitrogen fixation have been also incorporated. The database contained a total of 111,502 sequences and also a finish list of your genes included during the database will be located in Extra file 2. Table S5.
The searches for the genes to incorporate inside the database with the NCBI website have been to your Nucleotide assortment within the International Nucleotide Se quence Database Collaboration with limits, which excluded sequence tagged web sites, third get together annotation sequences, substantial throughput genomic sequences, patents, and full genome shotgun sequences. Additional limits were the search our website discipline was gene name along with the molecule was genomic DNA RNA, We also excluded hits that included finish genome in any discipline. The nearby BLASTN was performed at Situation Western Reserve Universitys Genome and Transcriptome Evaluation Core facility. Quite a few sequences in our database were complete chromosome sequences that included genes besides the N metabolism genes we were considering.
If se quences in the metagenomes matched with these information base entries, they had been only retained should the gene region of the BLASTN match was to a N metabolic process gene of interest The BLASTN comparison integrated an e worth cutoff of read this article 10 5 or reduced and sequence similarity cutoff of 50 base pairs or better. Statistical analysis The Statistical Evaluation of Metagenomic Profiles program was used to examine the NO3 and N metage nomes by identifying the proportional representation of different metabolic or phylogenetic groups and determin ing when they had been statistically various among the two metagenomes with two sided Fisher exact exams, The MG RAST functional matches in any respect ranges and taxo nomic matches with the class degree and higher had been com pared with Fisher exact exams. Storeys false discovery fee method was applied on the Fisher exact exams as a a number of comparison test correction, leading to q values, that are the FDR equivalent of p values.
Self-assurance in tervals have been determined with the Newcome Wilson method at 0. 05. Statistically major options that had significantly less than 5 sequences or very low effect sizes had been eliminated from the evaluation. Furthermore, a two sided chi square check, with Yates 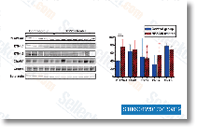 correction for continuity, was carried out, also applying STAMP, to the level two subsys tems. This test was completed specifically to investigate if any degree two EGTs from the N metabolism class have been statistically different which has a much less conservative check, Self-confidence intervals had been calculated and result size filters were made use of as with the Fisher actual tests.
correction for continuity, was carried out, also applying STAMP, to the level two subsys tems. This test was completed specifically to investigate if any degree two EGTs from the N metabolism class have been statistically different which has a much less conservative check, Self-confidence intervals had been calculated and result size filters were made use of as with the Fisher actual tests.
BRAF signal
The BRAF gene is a proto-oncogene
